New: K13 Molecular Surveyor supporting resistance surveillance
Monitoring the potential emergence, spread and prevalence of K13 mutations associated with resistant parasites in other regions can provide us with an early warning system to trigger rapid public health mobilisation to drive eradication. The WWARN team has developed an interactive tool that collates and summarises the prevalence of K13 mutations by geographical location and year.
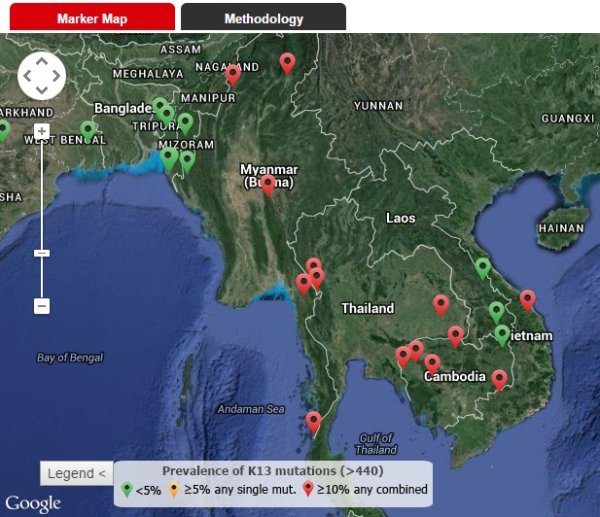
In the past, molecular markers of resistance to antimalarials have been identified long after the resistant parasites have emerged and spread widely throughout the world. With the advancements of molecular tracking and surveillance technology, we have the unusual opportunity to monitor the prevalence of mutations in the kelch gene (K13), associated with artemisinin resistance, as they evolve in time and space.
This early phase molecular information may allow us to support eradication or containment of the resistant parasites in the Greater Mekong Region, before they spread elsewhere. Monitoring the potential emergence, spread and prevalence of K13 mutations associated with resistant parasites in other regions can provide us with an early warning system to trigger rapid public health mobilisation to drive eradication.
To respond to this opportunity, the WWARN team has developed an interactive tool that collates and summarises the prevalence of K13 mutations by geographical location and year. The team has amassed data from over 5,400 samples from 11 published studies, covering sites in 22 countries, displayed over two maps - one specifically for Asia where the K13 markers are most prevalent, and another for the rest of the world.
“This tool is supporting surveillance efforts against further emergence and spread of artemisinin drug resistance from the Mekong Region of Southeast Asia to the rest of the world,” says Professor Carol Sibley, WWARN Scientific Director. “Researchers and policy makers can use this tool to view the temporal trends and prevalence of K13 mutations associated with resistance, and use this evidence to better inform key regional, national and international malaria monitoring strategies.”
Below the map there are other views of the same data that allow you to explore the temporal trends and compare two different sites of interest in more detail.
Dr. Venkatachalam Udhayakumar, Associate Director for Laboratory Science, Division of Parasitic Diseases and Malaria, Center for Global Health, commented: “It is extremely useful to have an easy point of access to the molecular information that is being generated on K13. The tool’s interactive format provides clear, standardized information that is easy to use and provides a useful asset for those seeking to better understand the implications of the prevalence of these molecular markers in the propeller region of the Kelch 13 gene.”
All of the information shown on the K13 Molecular Surveyor represents molecular analyses of K13 genotypes. It is not yet known whether the prevalence of certain mutations necessarily means that parasites are clinically resistant.
“The association of molecular markers with artemisinin resistance is an evolving field of research and therefore, as our understanding of these mutations grows and expands, the rules applied to the tool will evolve to reflect the latest research findings,” adds Dr Georgina Humphreys, WWARN Senior Data Manager. “This is an exciting time to be a researcher working in antimalarial resistance. We now have the tools and the technology to inform strategies to prevent further spread of resistance through molecular tracking. The K13 Surveyor complements current research approaches by giving us a useful overview of the situation of antimalarial resistance in Asia and beyond.”
Two other Molecular Surveyors display resistance markers for sulphadoxine pyrimethamine found on genes, dhfr and dhps; and resistance markers for Amodiaquine (AQ), Chloroquine (CQ), and Lumefantrine (LUM) found in P.falciparum pfmdr1 and pfcrt genes.
Have you read the latest paper on the prevalence of artemisinin resistant malaria across Myanmar?
Related publications:
Tun et al. Spread of artemisinin-resistant Plasmodium falciparum in Myanmar: a cross-sectional survey of the K13 molecular marker. Lancet Infectious Diseases. Published online February 20 2015. DOI: 10.1016/S1473-3099(15)70032-0.
