Approved uses of platform data
The following research applications for access to data on the Ebola Data Platform have been approved by the Platform’s independent Data Access Committee. The committee is overseen by the World Health Organisation and TDR, the Special Programme for Research and Training in Tropical Diseases. Researchers can apply to access data for research addressing key knowledge gaps concerning Ebola Virus Disease (EVD).
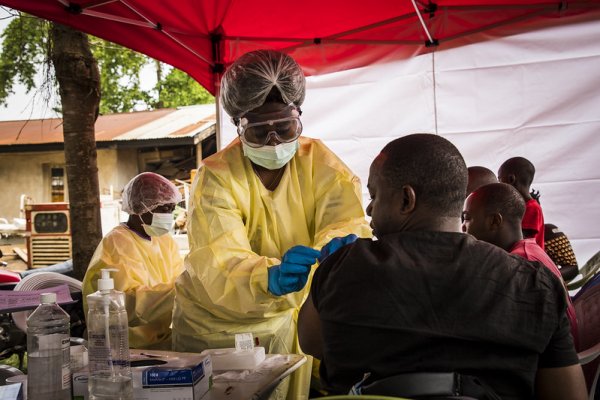
Approved applications
University Hospital Heidelberg, Germany
The Ebola virus disease (EVD) is an extremely infectious disease with a high case fatality rate, ranging from 25-90%. There is currently no known treatment that effectively neutralizes the virus. Instead, supportive treatments, such as oral rehydration solutions and interventions like infection prevention and control, and contact tracing, are the most crucial strategies to infection and disease management. This research uses the Ebola Data Platform’s compiled data from the 2014-2016 Ebola outbreak in West Africa to explore the determinants and predictors of mortality in EVD patients. This could include but is not limited to time to diagnosis and treatment, and host, viral and supportive care factors. Understanding the relationships between these factors could improve clinical management strategies, inform affected countries throughout the region, and improve the likelihood of survival should future outbreaks occur.
Brown University, USA
Although children represent a small number of the cases in many outbreaks of Ebola Virus Disease (EVD), the disease case fatality is high, especially in children less than five years old. Creating a predictive model for diagnosis for paediatric patients can help to rapidly identify patients who need care. Additionally, a predictive prognostic model for paediatric mortality allows clinicians to allocate resources appropriately. There are few studies to date that address diagnosis and prognosis of EVD in paediatric patients. This study will use retrospective data collected on paediatric patients (<18 years old) for whom outcome data is available in the Ebola Data Platform. Objectives of this research are to derive and externally validate a new diagnostic prediction model for paediatric EVD, and to derive and externally validate a new prognostic prediction model for paediatric EVD.
National Public Health Institute of Liberia
The general aim of this research is to improve the EVD patient management in the context of future outbreaks, addressing EVD research priorities and translating the research outcomes into concrete health policy. Specifically, the research will expand on the existing information on clinical management strategies with focus relating to the host, viral and supportive care factors associated with mortality. Efforts will be tailored toward survival analysis. Considering the patient parameters, the research analysis will undertake the effort to explore the relationships between changes in parameters and the associated risk of death. In the latter stages of the research project, a propensity score-matching approach will be used to re-evaluate the association between intervention and mortality to assess possible causal association. Findings from this study will be analysed epidemiologically and disclosed in a formal publication followed by stakeholder engagement in the affected countries.
Ministry of Health and Sanitation, Sierra Leone
The previous case definitions for EVD were not sufficient to identify all cases with signs and symptoms of Ebola which may be similar to other diseases like malaria and typhoid fever. This application seeks to improve on the case definition of EVD by looking at the huge amount of data emanating from the 2014-2016 Ebola outbreak in West Africa. The identification of the EVD-positive patient will help in providing specialist care, while ensuring that non-EVD patients continue with routine outpatient consultation. This would go a long way towards breaking the transmission of the disease and providing surveillance teams with adequate information on contact tracing and line listing. The study will look at key variables including signs and symptoms at presentation, history of exposure to EVD patients, laboratory results on admission and follow up, and pregnancy status. This project will put forward a more definitive case definition to inform policymaking and leverage the data to predict transmission trends and how onward transmission can be halted.
Gamal Abdel Nasser University of Conakry, Guinea / African Institute for Mathematical Sciences, Cameroon
This application seeks to characterise the evolution of the signs and symptoms of EVD using individual patient data (IPD) hosted by the Ebola Data Platform. In an emergency outbreak situation, it is important to have a reliable indicator for medical screening of patients and monitoring for evolution of signs and symptoms associated with EVD to identify those at increased risk of death, helping to optimise health care delivery. The study uses IPD meta-analysis, considered the gold-standard approach for synthesis of results across studies, where the volume of available data provide a unique opportunity to characterise the evolution of EVD symptoms. Clinical signs and symptoms at baseline and during hospitalisation period in Ebola treatment centre (ETC) patients with confirmed EVD are described, with longitudinal mixed effects modelling to characterise the population-averaged trajectory of the clinical signs and symptoms, and competing risk survival analysis to estimate the cumulative incidence of fever clearance, haematological recovery, and viral clearance.
Read the thesis (Amoa-Dadzeasah)
Read the thesis (Lemukong Ngufor)
École Polytechnique Fédérale de Lausanne (EPFL), Switzerland
The study will create an open-source collaborative data analysis platform that allows users to crowdsource diverse clinical insights on EVD from rapidly accumulating data across fragmented datasets, whilst preserving patient privacy and local ownership. Using the Ebola Data Platform to offer evidence on the applicability and performance of new, decentralised machine-learning methods, the study will create a platform for robust real-time data analysis during an outbreak. These methods will enable clinical researchers to generate reliable models using patient data that is distributed across sites, without the need for data owners to share any original data or provide their patient records to a centralised repository. This will strengthen patient privacy and incentivise collaboration and interoperability between competing research parties, which is especially relevant during health emergencies and in rural settings without reliable centralised medical infrastructure.
Read the preprint (Roschewitz)
Check out the learning platform
National Institute of Allergy and Infectious Diseases, U.S. National Institute of Health
During the West Africa Ebola outbreak in 2013-2016, clinical studies to evaluate potential therapies were mounted by separate organisations. The debate about how to evaluate effectiveness was considerable, especially around randomisation given the high mortality rate. In an effort to evaluate whether a model of mortality rates under “standard of care” could be created as a benchmark for putative therapies, a meta-analysis was conducted of Ebola studies carried out during the outbreak. Data from previous outbreaks were reviewed alongside the meta-analysis of the published studies from the 2013-2016 epidemic. The study evaluated how much information could be gleaned from the control groups of those studies and address the assumptions necessary for use in future outbreaks. A second objective was to evaluate whether the meta-analysis adds to the understanding of the potential efficacy of studied treatments.
Associated research publication
